SuperMogadon is a highly flexible search and sort tool that allows you to easily compare the client’s genotype with results from Genome Wide Association Studies* (GWAS) through the Opus 23 Pro database.
As you can see there are over 50+ pages of GWAS data in SuperMogadon, which would make grinding through the data rather impractical. Like most data displays in Opus 23 that deal with large amounts of data, SuperMogadon features a ‘filterable’ display. Type full or partial search terms into the search box at the upper left hand corner and SuperMogadon immediately displays only those results.
Click on the graph icon to display SNP distribution for that pathology or trait as a Manhattan Plot. Click on any column title to sort by that column.
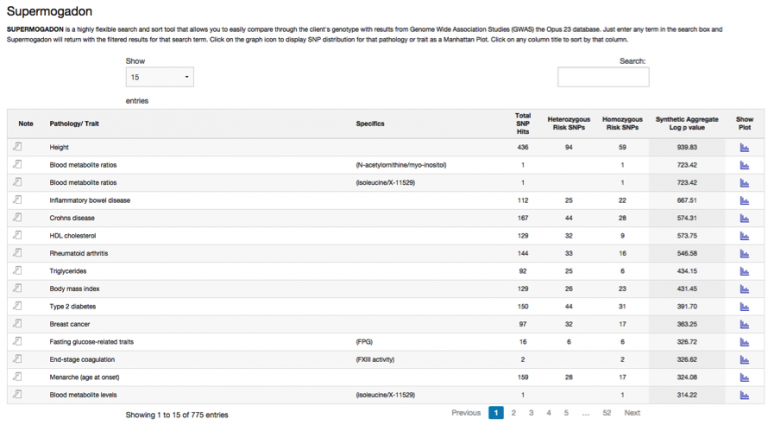
Clicking on the blue graph icon in the ‘Show Plot’ column will launch the SuperMogadon Manhattan plotter for that disease/trait.
TheSuperMogadon Viewer is a GWAS Manhattan plotter, a type of scatter chart used to display data with a large number of data-points. Genomic SNP coordinates (marked by chromosome) are displayed along the X-axis, with the negative logarithm of association P-value for the disease or pathology displayed on the Y-axis. Because the strongest associations have the smallest P-values (e.g., 10 −15), their negative logarithms will be the greatest (e.g., 15). In the example above, we see the graph for ‘Type II diabetes’ as a GWAS Manhattan plot.
Client SNP genotypes results are shape and color-coded:
- Gray-colored points denote client SNPs that do not contain the risk allele
- Orange-colored points denote that the client is homozygous for the risk allele
- Yellow-colored points denote that the client is heterozygous for the risk allele
- Square-shaped points signify that the SNP is in the GWAS and Opus 23 Pro databases and when clicked will trigger an information pop-up
- Circle-shaped points signify that the SNP is not in the Opus 23 Pro database but is in the GWAS database and is reported by 23andMe. When clicked these SNPs will bring up its GWAS PubMed reference article
Drag-select to zoom section or use the scroller at the bottom. Hover over any point to learn more. Clicking on any point triggers a full-information popup window. Like any other element in Opus 23 Pro, you can notate the SNP in SuperMogadon Viewer by clicking on the link to bring up the information pop-up, then clicking the ‘Add/Edit Note’ button at the top of the pop-up screen. You can also send any popup element directly to curation (so that it shows up in the Client Report.)
Opus 23 Pro subscribes to the philosophy of ‘TMTOWTDI’ (There’s more than one way to do it, pronounced ‘Tim Toady’.) The program was designed with this idea in mind, in that it ‘doesn’t try to tell the physician how to parse the data.’ Rather, it presents many different frameworks and cross-sections of the available client data, using a myriad of infographic treatments. This (and our ability as a species to excel at pattern recognition) dramatically increases the odds that a noteworthy finding will not go undiscovered.
* In genetic epidemiology, a genome-wide association study (GWA study, or GWAS), also known as whole genome association study (WGA study, or WGAS), is an examination of many common genetic variants in different individuals to see if any variant is associated with a trait. GWASs typically focus on associations between single-nucleotide polymorphisms (SNPs) and traits like major diseases.
